Combat against COVID-19 is still ongoing worldwide. Despite the great advancements in the realm of vaccine development, there is still a great need for efficient medications to treat the disease, especially in vulnerable populations (such as elderly people and subjects with background illnesses). A common strategy in drug development against COVID‑19 is inhibiting the binding of the viral Spike protein to the cellular ACE2 receptor, which facilitates the virus's entrance to the cells. While many attempts focus on blocking the viral Spike protein, this strategy is sensitive to viral mutations, making it less efficient. The teams of Profs. Schreiber and Rudich developed a stable protein that binds the ACE2 receptor at ultra‑high affinity (pM), effectively blocking viral infection without affecting the natural activity of the receptor. The protein can be administered through inhalation and can be combined with other potential anti-COVID-19 drugs such as interferon-β.
Combating the COVID-19 pandemic focuses on two main therapeutic strategies: developing efficient vaccines and identifying potent drugs. Despite the clear advantages of developing an effective vaccine to protect healthy populations, vulnerable populations (such as the elderly, people with diabetes, heart problems, and more) vaccination tends to be less effective. Moreover, it is unclear how long vaccination will protect from infection, and virus mutation may reduce vaccine effectiveness. Therefore, the development of an efficient COVID-19 drug is synergetic to vaccine development and of utmost importance.
The SARS-CoV-2 virus enters cells using its Spike protein, which binds to the ACE2 receptor expressed by cells throughout the respiratory system. Blocking the interaction between the SARS- CoV-2 Spike protein to the ACE2 receptor is a common strategy in developing drugs against COVID-19, with many attempts aiming at blocking the viral Spike protein, such as the development of monoclonal antibodies designed to target the Spike protein. However, commonly occurring mutations of the viral protein may reduce the effectiveness of such drugs. Alternatively, blockage of the ACE2 receptor may alter its enzymatic activity, critical for blood pressure homeostasis, thereby potentially causing side effects. Therefore, there is still an urgent, unmet need to develop anti-COVID-19 drugs that will effectively block virus entry to the cells.
To attenuate SARS-CoV-2 infection, the groups of Profs. Schreiber and Rudich constructed an ACE2 tight-binding protein inhibitor, which is stable, does not perturb the natural activity of the enzyme, and can be administered by inhalation.
The Schreiber laboratory developed novel yeast display methodologies that were used to select for an ultra-tight ACE2 binding protein, which inhibits the binding of SARS-CoV-2 to ACE2, thus preventing the virus from entering and infecting the host cell. The selection starting point was the receptor-binding domain (RBD) of the SARS-CoV-2 spike protein. Through the generation of multiple libraries (for selecting for protein-stability, binding affinity, rate of association, and protein expression) the teams generated the protein RBD-62, a 192 aa long (20Kd) variant of the RBD1. RBD-62 harbors multiple mutations, which enhance its binding to ACE2 as well as its stability. It has pM affinity, a 1000-fold improvement over the innate Spike protein binding (which it competes against). Using ACE2-expressing cells the teams have shown that RBD-62 outcompetes CoV-2 pseudo-virus and inhibits SARS-CoV-2 virus infection, with an EC50 of 6 nM. This, without inhibiting the activity of ACE2 in hydrolyzing angiotensin, which is an essential activity of ACE2 in regulating blood pressure and immune responses (Figure 1). The improved RBD-62 protein stability allowed for its nebulization at close to 100% yield, with the RBD-62-containing aerosol out-competing Spike protein binding. As a therapeutic agent, RBD-62 can be administered through inhalation on its own, or in combination with other drugs, for example, interferon-β. This should result in a synergetic effect of the drugs, as they affect orthogonal pathways. A significant advantage of our approach is that by blocking the endogenous ACE2 receptor, it is resistant against alterations of the virus, as it hampers the virus entry point to the cell, which is not affected by viral mutations.
- Treatment for COVID-19
- Stable and highly efficient
- Can be administered using inhalation
- Possible combination with other drugs used against COVID-19
- Resistant against alterations of the virus
- Does not affect the natural activity of its target, the ACE2 receptor
The teams have generated a small protein, RBD-62, with an optimized binding ACE2 and improved stability. They proved it successfully inhibits viral entry to cells in vitro as well as in the COVID-19 hamster model. They developed a protocol for an efficient nebulization of the protein, demonstrating it could be successfully combined with additional anti-COVID-19 drugs. Figure 2 provides an overview of the development plan, starting from RBD-ACE2 binding optimization to cell assays, development of an inhalation protocol, and animal experiments.
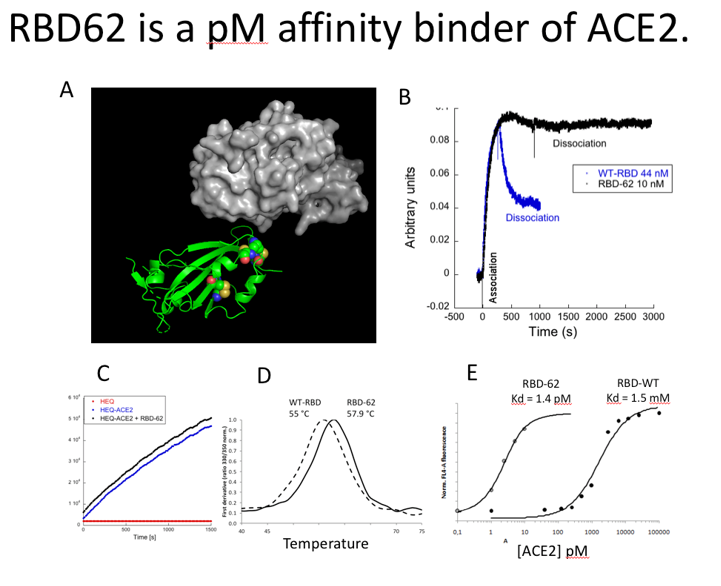
Figure 1. Comparing wild-type RBD to RBD-62. (A) shows the mutations Incorporated in RBD62 (ribbon) binding to ACE2 (gray surface). (B) biolayer interferometry technology implemented on an OCTET instrument with ACE2 immobilized on the surface. The binding constants were calculated using 3 different concentrations of RBD (of which a representative one is shown). (C) ACE2 enzymatic activity was determined on ACE2 expressing HEQ cells, showing no change in activity upon RBD-62 binding. (D) thermal denaturation of WT-RBD and RBD-62 as determined using a Tycho Nano-DSF. (E) binding affinity as measured using yeast display.
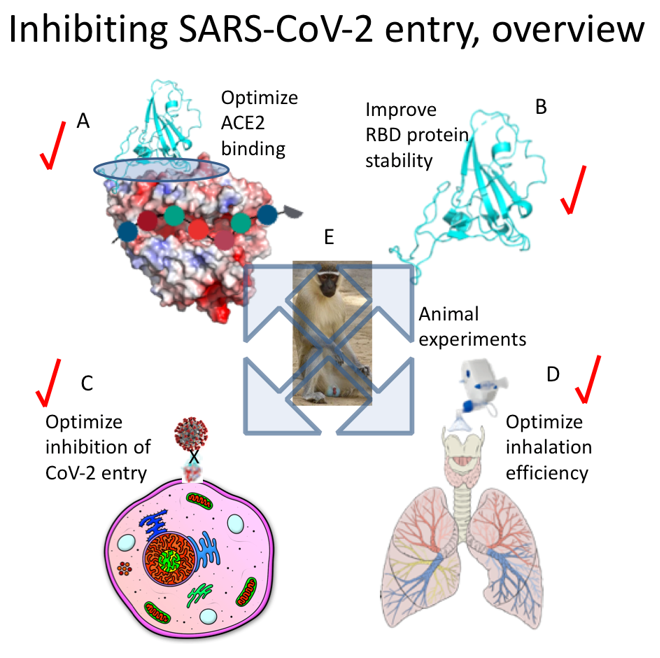
Figure 2. (A) optimizing binding of RBD to ACE2. (B) optimizing RBD protein stability. (C) determination of inhibition of viral entry using RBD. (D) development of protocols for RBD nebulization. (E) animal experiments to validate RBD efficiency in inhibiting SARS-CoV-2 infection. Tasks A-D were successfully completed.
Zahradník, Jiří, Shir Marciano, Maya Shemesh, Eyal Zoler, Daniel Harari, Jeanne Chiaravalli, Björn Meyer, et al. 2021. “SARS-CoV-2 Variant Prediction and Antiviral Drug Design Are Enabled by RBD in Vitro Evolution.” Nature Microbiology 6 (9): 1188–98. https://doi.org/10.1038/s41564-021-00954-4



