A bacteria (E. coli) engineered to fix atmospheric CO₂, converting it into organic biomass.
This breakthrough synthetic biology approach enables E. coli to use CO₂ as its sole carbon source by harnessing energy from formate, offering a potential solution to the major global challenge of reducing atmospheric CO₂ levels and a potentially unique platform for precision fermentation.
- Potential Precision Fermentation Platform: A modular platform for producing various chemicals with net negative CO₂ emissions by integrating synthetic metabolic pathways.
- Research Tool: Useful for studying and improving enzymes in the Calvin-Benson-Bassham (CBB) cycle, contributing to advancements in crop efficiency.
- Sustainable: Provides a way to sequester atmospheric CO₂
- Genetic Engineering Compatibility: E. coli-based standard laboratory techniques can be applied for genetic modification.
The Milo team has developed and characterized carbon-fixing strains of E. coli, demonstrating autotrophic growth through continuous laboratory evolution. These strains have been validated via carbon economy assessments and genomic sequencing to identify the mutations that support autotrophy.
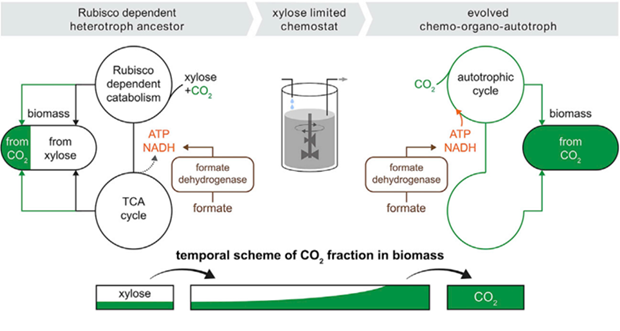
Experimental workflow: Engineered E. coli unable to grow autotrophically were cultured with limited xylose and excess formate/CO₂. Under this pressure, cells evolved to use CO₂ as the sole carbon source and formate for energy, leading to fully autotrophic clones that outcompeted the original strain.


